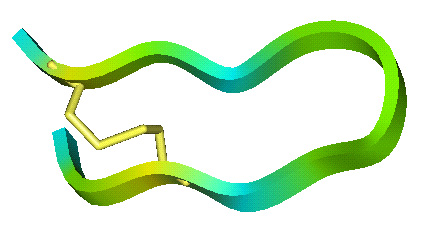Cationic Antimicrobial Peptides
In 1984, we proposed that cationic antimicrobial peptides were taken up by the self promoted uptake pathway. These peptides are ubiquitous in nature (more than 2000 are known) as important components of innate host defence mechanisms against pathogenic microbes.
We thus devised a recombinant DNA procedure for producing these peptides in bacteria and went on to investigate a wide variety of cationic antimicrobial peptides, and have published and patented peptides from all 4 structural classes.
Many variants have been made, most recently employing peptide arrays and machine learning techniques. As a result, structure-activity relationships have been established and structures were solved by 2D-NMR. The in vivo potential of these peptides as antibiotics and anti-endotoxins was demonstrated and the high throughput of peptide arrays permitted the identification of 8 amino acid peptides that had broad spectrum activity and was published in Nature, in addition to many mimetics.
A variety of patents derived from this work were licensed to four separate companies, the most advanced having been Migenix Inc, which reconfigured itself around this technology and demonstrated efficacy in phase III clinical trials in preventing lethal infections associated with indwelling catheters, while Cutanea has performed phase IIb trials showing efficacy in acne and Rosacea.
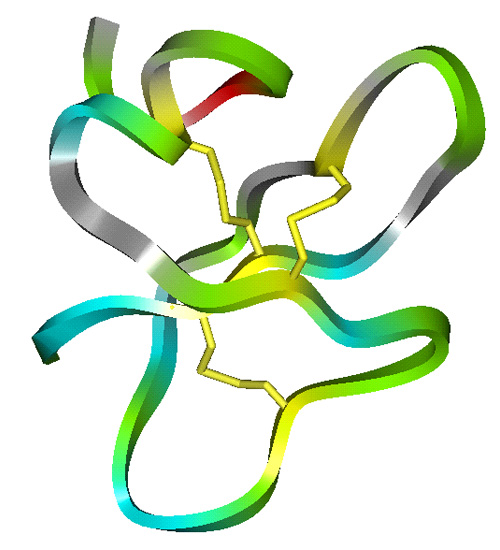
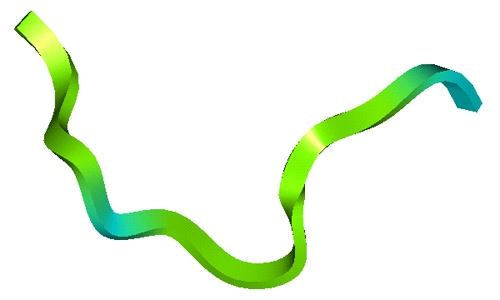
The lab has studied all four structural classes of cationic peptides:
- Amphipathic alpha-helices based on a fusion of silk moth cecropin and bee melittin;
- Extended peptides based loosely on the structure of the cattle neutrophil peptide indolicidin;
- Beta-sheet peptides including beta-hairpin peptides based on the horseshoe crab peptide polyphemusin, and cyclic beta-sheet peptides synthesized by our collaborator, Dr. Bob Hodges from the University of Alberta in Edmonton, based on the general structure of bacterial gramicidin S; and
- Loop peptides cyclized with a single cysteine disulphide based on cattle neutrophil bactenecin.
Structures of Cationic Peptide Types (Click to enlarge images)
We have studied the mechanism of action of representatives of most of these classes. Basically these peptides interact with the surface of Gram negative bacteria and are taken up by self-promoted uptake. They then insert into the cytoplasmic membrane under the influence of the transmembrane electrical potential gradient (which in bacteria is about -150 mV oriented internal negative so as to electrophorese the cationic peptides towards the membrane). They assemble in the membrane into multi-state channels which we have described via the “aggregate model”, and in many cases cross the cytoplasmic membrane to access cytoplasmic targets, or in some cases permeabilize the cytoplasmic membrane barrier. It should be stated that in the past many people in the antimicrobial peptide field favoured the latter mechanism for most peptides; however, our own published evidence appears to be more consistent with cytoplasmic targets for many peptides.
The better cationic peptides act very rapidly (within minutes) to kill cells and have very broad ability to kill microbes including the most important Gram negative and Gram positive pathogenic bacteria as well as fungi like Candida albicans. They are by and large unaffected by the most common clinical mechanisms of antibiotic resistance, and in our hands do not easily select resistant mutants even after multiple passages on sub-MIC doses of cationic peptides. We have been able to demonstrate that certain a-helical peptides are effective against systemic infections of mice by P. aeruginosa, and are also effective against chronic rat infections when delivered by aerosol.
We have evaluated different methods for MIC determination for cationic peptides and have proposed a standard method as described in our MIC methods paper (Wiegand, I., K. Hilpert, and R.E.W. Hancock. 2008. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nature Protocols 3:163-175).